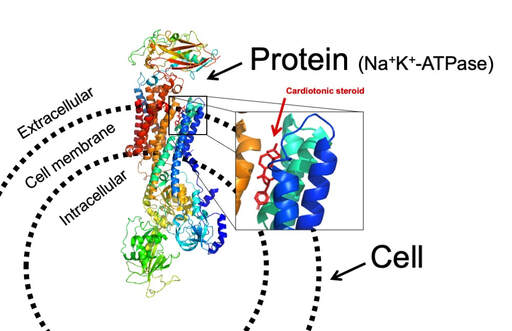
 Schematic diagram showing crystal structure of Na+K+-ATPase (PDB 4RES) with a bound cardiotonic steroid (bufalin).
Schematic diagram showing crystal structure of Na+K+-ATPase (PDB 4RES) with a bound cardiotonic steroid (bufalin).
Background
Cardiotonic steroids are common steroids that have been co-opted by many plants and animals for use as chemical defense. These steroids have one molecular target: the Na+K+ATPase, which is an important protein found in the membrane of all animal cells. By inhibiting these proteins, cardiotonic steroids can cause a heap of physiological irregularities, especially with regard to cardiac contractility. Despite this toxicity, many animals have evolved resistance to these steroids through target-site insensitivity of their Na+K+ATPases and can thus consume large quantities of cardiotonic steroids with no apparent ill effects.
Cardiotonic steroids are common steroids that have been co-opted by many plants and animals for use as chemical defense. These steroids have one molecular target: the Na+K+ATPase, which is an important protein found in the membrane of all animal cells. By inhibiting these proteins, cardiotonic steroids can cause a heap of physiological irregularities, especially with regard to cardiac contractility. Despite this toxicity, many animals have evolved resistance to these steroids through target-site insensitivity of their Na+K+ATPases and can thus consume large quantities of cardiotonic steroids with no apparent ill effects.
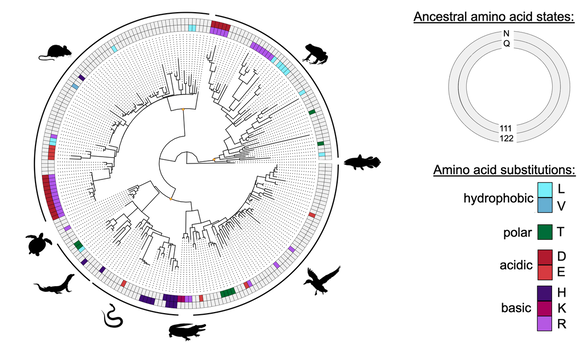 Amino acid substitutions at two key sites (111 and 122) on the protein have been repeatedly implicated in conferring this adaptation across diverse lineages.
Amino acid substitutions at two key sites (111 and 122) on the protein have been repeatedly implicated in conferring this adaptation across diverse lineages.
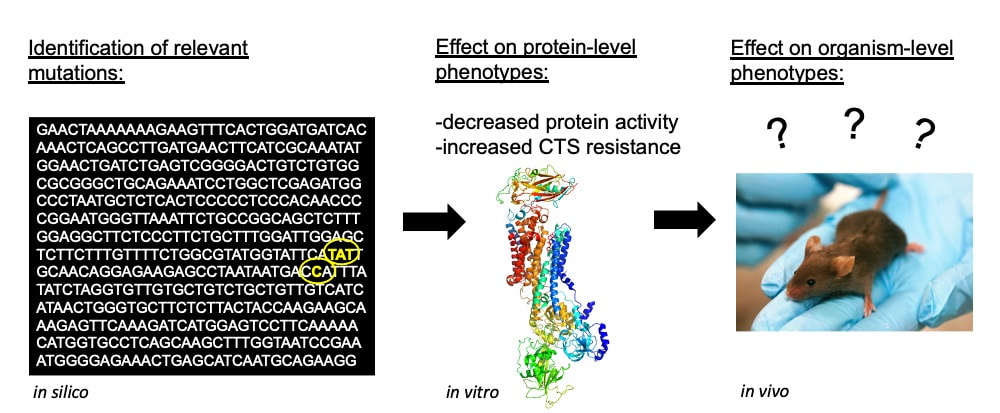
How predictable is the evolution of target-site insensitivity of Na+K+-ATPase (in silico)?
Our evolutionary analyses of the Na+K+ATPase gene family (ATP1A1--4) has revealed remarkable convergence in the molecular mechanisms by which many resistant animals have achieved this novel protein function. In all known cases, target-site insensitivity has been traced to a handful of amino acid substitutions at a key region of the ATP1A gene paralogs. By surveying the ATP1A gene family across tetrapods, we've compiled a rich body of comparative data that we're using to map the evolutionary landscape of this adaptation and assess the extent of parallelism, convergence, and divergence in resistance-conferring amino acid substitutions. See works published in Proceedings of the Royal Society B and PLoS Genetics.
Our evolutionary analyses of the Na+K+ATPase gene family (ATP1A1--4) has revealed remarkable convergence in the molecular mechanisms by which many resistant animals have achieved this novel protein function. In all known cases, target-site insensitivity has been traced to a handful of amino acid substitutions at a key region of the ATP1A gene paralogs. By surveying the ATP1A gene family across tetrapods, we've compiled a rich body of comparative data that we're using to map the evolutionary landscape of this adaptation and assess the extent of parallelism, convergence, and divergence in resistance-conferring amino acid substitutions. See works published in Proceedings of the Royal Society B and PLoS Genetics.
|
How accessible is the evolution of target-site insensitivity of Na+K+-ATPase (in vitro)?
In another layer of research, I investigate the molecular mechanisms that have constrained the evolution of cardiotonic steroid resistance across the animal kingdom. I combine bioinformatics with protein engineering and in vitro biochemical assays to functionally investigate the importance of ancestral starting points in the evolution of this adaptation. For example, we found that the phenotypic effects of resistance-conferring mutations depend strongly on the gene sequence in which they occur. Whereas one mutation would give resistance to the protein of one species, it would have a completely different effect in that of another. Upon close examination of a few case examples, we were able to show functionally that full adaptive transformation of the proteins was only possible when a combination of interactive mutations where present, not just those that confer resistance (see work published in Current Biology and Molecular Biology and Evolution. |
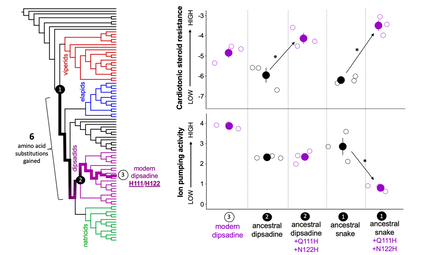
Empirical framework for understanding the importance of ancestral starting points in evolution. Phylogeny of snakes shows the evolution of the ancestral snake (node 1), to the ancestral dipsadine (node 2), to the modern dipsadine (node 3). Protein engineering and in vitro assays reveal that the modern dipsadine mutations (Q111H and N122H) produce resistance on both ancestral backgrounds. They have no effect on overall protein function (ion pumping activity) in the ancestral dipsadine background but cause a significant loss of function on the ancestral snake protein. Six amino acid substitutions were gained in the evolution of the ancestral snake NKA to the ancestral dipsadine NKA. Within these six are key changes that mitigate the negative pleiotropic effects of Q111H and N122H on protein activity and thus opened up the possibility of evolving these resistance-producing mutations in the dipsadines.
|
What effects does gaining cardiotonic steroid resistance have on other traits?
A lot of my early work in this system involved investigating the physiological implications of cardiotonic steroids resistance, particularly in snakes, which were an ideal vertebrate model because there are many resistant species that feed on cardiotonic steroid-defended toads. The culmination of this early work, resulted in the identification of many interesting physiological patterns. For example, we found that resistant snakes with a high percentage of toads in their diets had significantly enlarged adrenal glands (see work published in Journal of Zoology). We later found that these snakes produced high levels of circulating aldosterone, which is a steroid hormone produced and secreted by the adrenal glands and which signals the expression of Na+K+ATPases (see work published in Gene and General and Comparative Endocrinology).
While experiments that investigate the effects of genetic changes on proteins as an estimate of fitness have expanded our understanding of the mechanisms of evolution, the actual effect of such changes extends beyond protein function to whole organism physiology (i.e., gene → protein function → cell physiology → organismal physiology). For example, what we measure as an insignificant reduction in ion pumping function at the protein level might translate to significant physiological consequences at higher biological levels. To gain a comprehensive understanding of the mechanisms that mediated the evolution of an adaptation, we must test the causal links between specific evolutionary changes in gene sequence and the resulting changes in protein function, cell physiology, and eventually, whole-organism physiology.
As a first step towards addressing this gap, I'm currently establishing an in vitro system to test the effects of resistance-conferring mutations on different genetic backgrounds, as I have been doing, but with an additional layer of experiments at the cellular level. These experiments will allow us to monitor how the evolution of target-site insensitivity of Na+K+-ATPase altered cellular signaling and homeostasis. These experiments are also meant to provide a transitionary step on the way to establishing an in vivo system for testing these effects on whole organism physiology. With the establishment of both systems, we will have a full gene, to protein, to cell, to whole organism experimental model for investigating the dynamics of the adaptive process in the most comprehensive and direct way possible.
A lot of my early work in this system involved investigating the physiological implications of cardiotonic steroids resistance, particularly in snakes, which were an ideal vertebrate model because there are many resistant species that feed on cardiotonic steroid-defended toads. The culmination of this early work, resulted in the identification of many interesting physiological patterns. For example, we found that resistant snakes with a high percentage of toads in their diets had significantly enlarged adrenal glands (see work published in Journal of Zoology). We later found that these snakes produced high levels of circulating aldosterone, which is a steroid hormone produced and secreted by the adrenal glands and which signals the expression of Na+K+ATPases (see work published in Gene and General and Comparative Endocrinology).
While experiments that investigate the effects of genetic changes on proteins as an estimate of fitness have expanded our understanding of the mechanisms of evolution, the actual effect of such changes extends beyond protein function to whole organism physiology (i.e., gene → protein function → cell physiology → organismal physiology). For example, what we measure as an insignificant reduction in ion pumping function at the protein level might translate to significant physiological consequences at higher biological levels. To gain a comprehensive understanding of the mechanisms that mediated the evolution of an adaptation, we must test the causal links between specific evolutionary changes in gene sequence and the resulting changes in protein function, cell physiology, and eventually, whole-organism physiology.
As a first step towards addressing this gap, I'm currently establishing an in vitro system to test the effects of resistance-conferring mutations on different genetic backgrounds, as I have been doing, but with an additional layer of experiments at the cellular level. These experiments will allow us to monitor how the evolution of target-site insensitivity of Na+K+-ATPase altered cellular signaling and homeostasis. These experiments are also meant to provide a transitionary step on the way to establishing an in vivo system for testing these effects on whole organism physiology. With the establishment of both systems, we will have a full gene, to protein, to cell, to whole organism experimental model for investigating the dynamics of the adaptive process in the most comprehensive and direct way possible.
Proudly powered by Weebly