Battle between molecular evolutionary forces shapes toxin resistance in Neotropical grass frogs5/19/2021 Reporting on: Mohammadi S, Yang L, Harpak A, Herrera-Álvarez S, Rodríguez-Ordoñez MP, Peng J, Zhang K, Storz JF, Dobler S, Crawford AJ, Andolfatto P. 2021. Concerted evolution reveals co-adapted amino acid substitutions in Na+ K+-ATPase of frogs that prey on toxic toads. Current Biology. [link]  Neotropical grass frog, Leptodactylus insularium. Photo by Brian Gratwicke (2011). Neotropical grass frog, Leptodactylus insularium. Photo by Brian Gratwicke (2011). This month I wanted to write about a study I had the privilege of working on with a talented and diverse group of researchers from around the world. Our multidisciplinary teamwork allowed us to uncover a very cool story of evolution involving Neotropical grass frogs from South America. This story centers around gene duplication – a mutational event wherein an extra copy of a gene is produced. Gene duplication is a powerful tool in evolution because it can liberate one copy of the gene to mutate and produce new roles without causing significant detrimental effects in the organism—the extra copy can act as a sort of safety backup. We explored the evolution of a gene duplication that had been identified in Neotropical grass frogs (genus Leptodactylus) and consequently unraveled the remarkable pathway that led them to gain a specialized adaptation—toxin resistance. The gene in question is ATP1A1, which codes for a protein vital to all animal cells—the sodium-potassium pump (or Na+K+-ATPase). We surveyed genetic material from many species of Neotropical grass frogs. We found that the ATP1A1 duplication was present in all species of the genus. We further found that there are 12 shared amino acid differences between the two gene copies. Two of these amino acid substitutions were known in other animals to provide resistance to common toxins known as cardiotonic steroids. These toxins are used as a chemical defense in toads, insects, snakes, and many plants. They target and disable sodium-potassium pumps. Because Neotropical grass frogs are known to feed on toads, having a resistant copy of this gene makes sense. This was a textbook example of neo-functionalization, wherein a gene acquires a new function after a duplication event. In this case, a new function evolved between a resistance-conferring copy, which we dubbed the R copy, and a copy that retained ancestral susceptibility, which we dubbed the S copy To trace the origins of this duplication we constructed a genealogy of the ATP1A1 gene using DNA sequences. However, this genealogy showed that the gene duplication occurred multiple times, once in each species, and was then followed by 12 parallel amino acid substitutions. This didn’t make sense considering how unlikely it is for the identical duplication and subsequent identical mutations to occur multiple times. The true origin of the duplication and subsequent 12 substitutions was revealed when we constructed our genealogy using amino acid sequences (i.e., protein sequences): they occurred once in the shared common ancestor of all Neotropical grass frogs and were subsequently inherited by all the descendent species. Although the DNA-based genealogy didn’t show us the origins of the gene duplication, it exposed another hidden story. The DNA patterns show that the R and S genes remained very similar to each other throughout tens of millions of years because of frequent non-allelic gene conversion (NAGC). NAGC is a molecular mechanism that homogenizes differences between two gene copies. This is known as the nefarious concerted evolution, which can stop neo-functionalization by working to keep duplicated genes identical. Frequent NAGC can result in a genealogy in which duplicated genes (two different genes) in the same species appear to share a more recent common ancestor than with their counterparts (the same genes) in other species. This explains why our DNA-based genealogy showed multiple origins of the gene duplication. In comes the hero, natural selection, which can oppose the homogenizing force of NAGC by promoting divergence between the duplicated genes. Hidden in the ATP1A1 gene sequences are short segments of divergence between the duplicates, which are strongholds that evaded concerted evolution. There are 12 of them, and they strongly distinguish R and S. We ran a computational model to determine how strong natural selection would have had to be to overcome the forces of concerted evolution at these 12 sites. The results suggested that substantial selection had to act to create such strongholds of “escape” from the hold of frequent NAGC. The fact that strong selection maintained these 12 amino acid substitutions implies that they’re functionally important and collectively contribute to organismal fitness. We knew that 2 of these 12 substitutions were associated with cardiotonic steroid resistance, but we had no clue what the other 10 were doing. To figure this out, we performed protein engineering experiments and functional assays. We produced the S and R copy proteins in the lab, and then added various combinations of the 12 R substitutions onto the S copy gene to measure their effects. Our results confirmed that the 2 resistance-conferring substitutions were indeed responsible for resistance in the R protein, while the additional 10 did nothing to enhance resistance. However, our results revealed that the 2 resistance-conferring substitutions come at a high price. When they’re added both individually and together on the S protein, they drastically reduce the protein’s ability to function. Thus, the resistance-conferring substitutions provide an adaptive advantage at the expense of protein function. This is where the functional importance of the other 10 substitutions is revealed. When these 10 substitutions are added together with the 2 resistance-conferring ones, protein function is rescued. Thanks to the genetic signals left behind from a tension between molecular evolutionary forces (concerted evolution vs. natural selection), we were able to trace the functional changes underlying this incredible adaptation in frogs. The multidisciplinary talents of our global research team were essential to unravel this story, and highlight how diverse collaborations can better unravel biological stories. by Shabnam Mohammadi
3 Comments
Reporting on: Haiser HJ, Gootenberg DB, Chatman K, Sirasani G, Balskus EP, Turnbaugh PJ. 2013. Predicting and manipulating cardiac drug inactivation by the human gut bacterium Eggerthella lenta. Science. 341:295–298. [link]  Every year we learn more about the different ways our gut microbiota influences our health and behavior. This microscopic community can change our appetites, cravings, immune functions, moods, and more. One could even argue that we’re just flesh robots controlled by microbes. Perhaps less commonly known is their ability to alter your resistance to certain drugs and toxins. Let’s discuss how this works using digoxin as a case example. Digoxin is a highly effective drug for treating many heart conditions. It’s isolated from the foxglove plant (Digitalis lanata) and belongs to a group of compounds known as cardiotonic steroids. Cardiotonic steroids come in two varieties—the plant-derived cardenolides, such as digoxin, and the animal-derived bufadienolides. Cardenolides are produced by many plants for chemical defense against insect hosts and herbivores. Similarly, bufadienolides are produced by toads and fireflies for chemical defense against predators. Surprisingly, cardiotonic steroids are also produced in much lower concentrations in our own bodies, where they serve as a signaling hormone [1]. All in all, this highly versatile group of compounds can play the role of poison, drug, or hormone.  The foxglove plant (Digitalis lanata), common throughout Europe, western Asia, and northwestern Africa. The foxglove plant (Digitalis lanata), common throughout Europe, western Asia, and northwestern Africa. All cardiotonic steroids do the same thing—find and disable proteins known as sodium-potassium pumps. These proteins exist in our cell membranes and, as their name suggests, they pump sodium and potassium in and out of cells. They’re absolutely vital for the maintenance of many cellular and physiological systems and their disablement, depending on the extent and context, can be either therapeutic, neutral, or lethal. Here’s how that works: when sodium-potassium pumps are disabled, they stop transporting sodium out of the cell, causing a sodium overload inside the cell. To compensate for this, the cell activates a protein that takes sodium out of the cell in exchange for calcium. This solves the sodium problem, but causes a calcium excess. To fix this, the cell pushes all that excess calcium into an organelle called the sarcoplasmic reticulum, which then delivers it to muscle cells. Notably, small amounts of calcium delivered to muscles trigger contractions. In this case, the excess calcium triggers strong and prolonged contractions [2]. If you ingest a toxic dose of cardiotonic steroids (e.g., eat one of the plants or animals mentioned above), death will likely ensue as a result of cardiac arrest—imagine the heart muscles contracting erratically and seizing up. That being said, for someone fighting heart failure, wherein the heart muscles are too weak to pump blood efficiently, cardiotonic steroids can have a therapeutic effect at the right dose. In fact, cardiotonic steroids have been used in this capacity for over 3000 years. The oldest records have been traced all the way back to ancient Egypt, where the medical Ebers Papyrus (1555 BC) documented the use of plants containing cardiotonic steroids to treat cardiac conditions [3]. Today, their therapeutic potentials are so vast they’ve even spread to the realm of male birth control [4] and anticancer drugs [5].  Vincent van Gogh's "Portrait of Dr. Gachet" features a familiar plant in the foreground. Vincent van Gogh's "Portrait of Dr. Gachet" features a familiar plant in the foreground. On a slightly tangential but fun note, there’s an interesting connection between these compounds and Vincent van Gogh. Van Gogh is widely believed to have consumed digoxin from the foxglove plant, either recreationally or for therapeutic reasons. You often find this plant in his paintings, like "Portrait of Dr. Gachet." Among the side effects associated with digoxin, there are three that stand out when you consider van Gogh’s distinct painting style: disturbed color vision, seeing halos around lights, and the overbright appearance of lights. These attributes are particularly conspicuous in some of his works, like the famous "Starry Night." Whether digoxin really did contribute to van Gogh’s paintings, however, remains a mystery. Disclaimer—please don’t eat poisonous plants to improve your painting. Despite all the therapeutic potential, cardiotonic steroids are difficult to prescribe—they have a narrow therapeutic dose range and reactions to the drugs vary widely on an individual basis. In the U.S. alone, thousands of cases of cardiotonic steroid poisoning are reported each year, many of which occur at medical centers [6]. Here’s where gut microbes come in. The human gut microbiota is well known to alter the activity and toxicity of drugs in the intestines. By doing so, they can directly change how much of the original drug is then circulated in the blood, and ultimately delivered to the drug target. Breakthrough work published in Science back in 2013 [7] by Drs. Henry Haiser and Peter Turnbaugh and colleagues at Harvard University has revealed the incredible way that one species of bacteria in your gut can protect you from the toxic effects of digoxin. First, a little history. Back in the 1980’s, clinical trials investigating how the human body gets rid of orally ingested digoxin found that some individuals who exhibited lower levels of blood digoxin, post-dose, excreted an inactive form of the compound in their stool. To investigate further, scientists grew bacteria from the stool of these individuals and found that they could metabolize digoxin into an inactive form. Conversely, bacteria from stool cultures of individuals that didn’t excrete inactive digoxin could not metabolize the drug. Further confirming the source of inactivation to gut microbes, scientists found that individuals who excreted inactive digoxin stopped doing so after being given antibiotics [8]. Several years later, the source was further pinpointed to one species of Actinobacterium known as Eggerthella lenta (E. lenta). In come Haiser and colleagues in 2013 with a multi-angle approach aimed at figuring out exactly how these bacteria inactivate digoxin. By growing E. lenta under two conditions—one with and one without digoxin—then measuring how the bacteria’s gene expression levels changed under these treatments, they found that one set of genes was significantly upregulated under exposure to digoxin. They refer to these as cardiac glycoside reductase (cgr) genes. To confirm that these genes were indeed responsible for digoxin inactivation, they tested the digoxin inactivation capacities of three strains of E. lenta, two of which lacked cgr genes. Indeed, only the strain with the cgr genes were able to inactivate/metabolize digoxin. This means that you can’t predict a person's ability to resist digoxin by simply testing for the presence or absence of E. lenta in their gut—the specific strain matters. To further test whether E. lenta's cgr genes were responsible for a person’s ability to metabolize digoxin, Haiser and colleagues measured the density of E. lenta, the level of cgr genes expressed, and the digoxin inactivation capacity of the stool microbes of 20 unrelated individuals. They found that individuals with high levels of cgr gene expression in their gut microbiota had the highest levels of digoxin inactivation. Even more to the point, they found that a person’s level of cgr expression was a much better predictor of their ability to metabolize digoxin than their level of E. lenta. Being that biological mechanisms typically involve complex interactions, Haiser and colleagues tested whether other factors can significantly influence cgr-mediated digoxin inactivation. Previous studies had shown that the amino acid arginine somehow suppresses E. lenta’s ability to metabolize digoxin. Amino acids are what proteins are made of, and accordingly, common dietary sources of arginine include anything with protein, such as meats, dairy, eggs, and seeds. Haiser and colleagues decided to include this factor in their tests and grew bacteria under two treatments—with and without arginine—in addition to with and without digoxin. They found that although arginine promoted stronger growth in E. lenta, it suppressed the expression of cgr genes, leading to a reduction in their inability to inactivate digoxin. The influence of arginine led Haiser and colleagues to hypothesize that the amount of protein in an individual's diet could affect their ability to metabolize digoxin. They tested this hypothesis on germ-free mice—mice with zero bacteria in their digestive systems. They colonized the mice with E. lenta, then fed one group a low protein diet and the other a high protein diet. They then dosed the mice with digoxin and measured how much made it to their blood, urine, and stool. Their work revealed that indeed, mice fed a high protein diet showed significantly lower levels of inactivated digoxin in their stool and, consequently, higher blood and urine digoxin than the mice fed a low protein diet. Another interesting finding was that the level of digoxin inactivation increased when E. lenta were co-cultured with other gut bacteria. An explanation for this synergy could be that E. lenta simply grows better when factors such as pH, nutrient concentration, and temperature are altered by other bacteria. Another reason could be that reduction of arginine availability caused by increased competition for arginine by other bacteria reduces cgr gene suppression. Overall, the work of Haiser and colleagues made a big step forward in our understanding of the mechanism through which gut microbes alter digoxin. Since then, other studies have expanded on this work by investigating more E. lenta strains and close relatives. We’ve now learned that more genes make up the cgr cluster than previously thought, and that the cgr-mediated digoxin inactivation mechanism generally works for all plant-derived cardiotonic steroids (i.e., cardenolides), though not the animal-derived ones [9]. All together, these studies are bringing to light the importance of considering the composition and function of our bodies’ microbial communities and how they interact with each other when developing pharmacological treatments, and ultimately, advancing our ability to properly prescribe medication. by Shabnam Mohammadi
Extreme radiation resistance achieved by mutations in DNA repair mechanisms of black yeasts.1/5/2021 Reporting on: Romsdahl J, Schultzhaus Z, Chen A, Liu J, Ewing A, Hervey J, Wang Z. 2020. Adaptive evolution of a melanized fungus reveals robust augmentation of radiation resistance by abrogating non‐homologous end‐joining. Environmental Microbiology. [link]  Ramsar, Iran. Source: Wikimedia Commons. Ramsar, Iran. Source: Wikimedia Commons. It goes without saying that ionizing radiation can be a scary thing. This “invisible force" can dismantle you from within by breaking your DNA to pieces. You’d need a hefty dose to suffer such consequences, though. In fact, we’re exposed to relatively harmless levels of background radiation regularly. For example, residents of Ramsar, Iran—the city that holds the current record for highest background radiation—are exposed to an average dose of 10 Gy per year [1]. (The Gray (Gy) is a dose unit that reflects the amount of energy delivered to a mass of tissue.) To give you some perspective, in the aftermath of the 1986 Chernobyl disaster, evacuees were reported to have received a whole-body dose of 0.002 to 0.66 Gy [2]. Ramsar's natural ionizing radiation comes from its geology, which is dotted by hot springs that move radioactive decay to the earth’s surface. Surprisingly, there’s been no increase in rates of cancer, leukemia, or other radiation-associated health complications among the residents [1]. Some researchers even argue that low to moderate exposure to ionizing radiation can have a therapeutic effect by activating DNA repair systems and other protective mechanisms in our bodies—not far off from how vaccines work [3]. However, after a certain threshold, there’s no escaping the damage that ionization radiation can inflict. The average acute dose that would cause radiation sickness is 0.7-10 Gy [4]. No human is expected to survive an exposure higher than 10 Gy, and at 50 Gy, death is expected within three days [4]. Keep in mind that these are acute thresholds—the same doses spread over a longer time may not be so damaging because the body will have time to repair between exposures. Unlike non-ionizing radiation (e.g., microwaves and UVA light), which only has enough energy to produce heat, ionizing radiation can break atomic bonds (i.e., damage to the body at the molecular level) and it’s penetrative, so damage can be inflicted internally. There are four types of ionizing radiation: alpha (α), beta (β), neutrons, and electromagnetic waves such as gamma (γ) rays. Alpha radiation particles are the heaviest and are emitted by naturally occurring radioactive materials like uranium and radon (e.g., in smoke detectors). Because they’re so heavy, they can’t penetrate barriers very well-- they can’t even get through paper (although inhalation can pose problems). Beta particles consist of single electrons and are emitted by radioactive isotopes of elements like hydrogen (e.g., tritium) and carbon (e.g., carbon-14). Beta particles don’t penetrate very well either and can be stopped by a layer of clothing. Neutrons are a form of ionizing radiation that consist of, well, free neutrons. They’re commonly produced from the splitting of atoms in nuclear reactors. Neutrons can travel a long way and can penetrate most barriers, even lead. The only way to stop them is with large quantities of water or other materials made of very light atoms, such as boron, which can bounce and absorb them (concrete is often used in this capacity). Finally, we have electromagnetic waves, like X-rays and γ rays, which are commonly used in medicine because they can penetrate our bodies. They’re produced when a particle, like an electron, is accelerated by an electric field or by the radioactive decay of atomic nuclei. 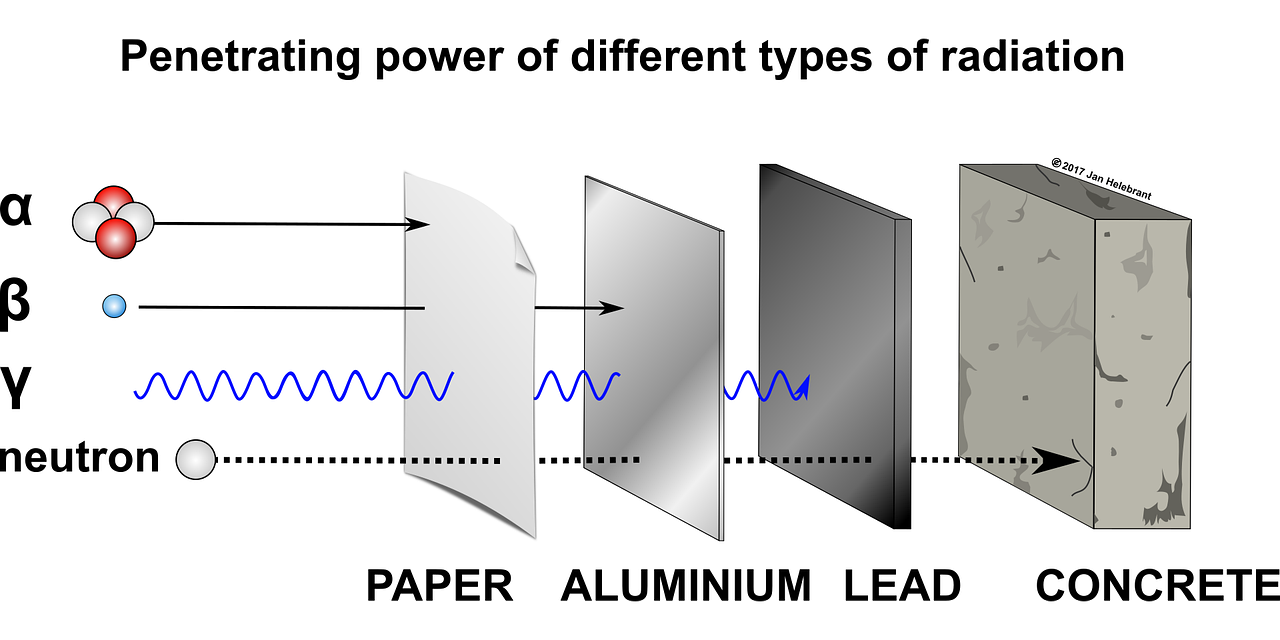 Penetration capacities of ionizing radiation. Source: Image by Jan Helebrant from Pixabay.  Species of black yeast known as Exophiala phaeomuriformis. Source: Wikimedia Commons. Species of black yeast known as Exophiala phaeomuriformis. Source: Wikimedia Commons. Now that we have a basic understanding of ionizing radiation, let’s talk about… yeast. Fungi are among the most radiation resistant eukaryotic organisms on Earth. Black yeasts in particular have been found in highly radioactive environments, such as the cooling pools of nuclear reactors, the stratosphere, and the damaged nuclear reactor at Chernobyl [5]. The remarkable resistance of black yeast to ionizing radiation presents an opportunity for scientists to pinpoint the genetic and biochemical underpinnings of this adaptation and consequently inform efforts aimed at developing therapies and safeguards against ionizing radiation. This can be of particular use to solving a major limitation of human space exploration: long-term exposure to cosmic ionizing radiation imposed by the space environment [6]. Several studies have attempted to uncover the genetic underpinnings of radiation resistance by searching for adaptive mutations in the genomes (i.e., the full DNA content) of resistant organisms [7]. This basically involves comparing the genetic code of a highly resistant species to that of a closely related non-resistant species, looking for differences (e.g., mutations), then investigating those differences. For example, do they occur in genes that code for enzymes involved in DNA repair? Unfortunately, these studies have been more or less inconclusive due to the sheer expanse of genomic data and the complexity of how mutations facilitate novel abilities. Taking this a step further, some researchers have attempted to artificially produce radiation resistance in bacteria. Essentially, this involves blasting bacteria with radiation, selecting and growing the survivors, then repeating this over many generations until a highly resistant strain emerges. This method is known as "directed laboratory evolution" and through such experiments, scientists have repeatedly found that changes in genes coding for DNA repair proteins facilitated the evolution of radiation resistance [8, 9, 10]. In other words, bacteria "artificially" evolved resistance to ionizing radiation by improving their DNA repair mechanisms. In October 2020, Drs. Jillian Romsdahl and Zheng Wang of the Naval Research Laboratory in Washington, DC, and colleagues published the results of a directed evolution study involving black yeast, which revealed a surprising adaptive mechanism against radiation. After repeated rounds of artificial selection, Romsdahl et al. produced several strains of extremely radiation resistant black yeast. Following genetic examination and functional experimentation, they discovered that these highly resistant strains owed a portion of their resistance to the deactivation of one of their DNA repair mechanisms. That may seem completely counterintuitive, but hear me out because this is really cool… Here’s how they did it: black yeast, which already have resistance to ionizing radiation were subjected to extremely high doses of γ radiation. The survivors were grown and then subjected to another round of γ radiation. This process was repeated 15 times, and the amount of radiation they were subjected to was increased every five rounds. In the first five rounds, they were given a 4500 Gy dose of ionizing radiation (remember the lethal dose for a human is anything higher than 10 Gy!). In the next five rounds the survivors received 5000 Gy, and in the final five, 5500 Gy. The average survival rate in each round was 1%. The yeast were essentially directed to evolve even more resistance than they already had. Since DNA damage is accepted as the main cause of death from ionizing radiation, Romsdahl et al. proceeded to subject their newly evolved strains with all sorts of DNA damaging methods, including exposure to desiccation, UVC light, the carcinogen methyl methanesulfonate, and anti-cancer drugs like Bleomycin and Hydroxyurea. They found that although their newly evolved strains had acquired higher resistance to ionizing radiation, for several strains this came at a cost to their ability to repair other types of DNA damage and to their general fitness. Next, Romsdahl et al. sequenced and analyzed the full genomes of all the evolved strains to identify possible causative mutations—they found many. By examining the functional significance of these mutations, one clear pattern emerged: the repeated occurrence of detrimental mutations (i.e., frame shifts or premature stop codon gains) in genes involved in a DNA repair method known as “non-homologous end joining” (NHEJ), basically rendering the mechanism nonfunctional. This is surprising considering that NHEJ repairs DNA damage and you’d think you'd want that up and running when you’re blasted with ionizing radiation. Well, not exactly… To investigate this puzzling outcome further, Romsdahl et al. took fresh black yeast strains that had been genetically modified to not have certain genes associated with NHEJ (i.e., they didn’t have functioning NHEJ), and exposed them to 4000 and 6000 Gy of γ-radiation. Lo and behold, they found that the deletion of these genes resulted in increased resistance to γ-radiation. However, the most resistant of the evolved strains still demonstrated higher resistance than the non-evolved, genetically modified strains. This means that the genetic underpinnings contributing to the evolved strains’ resistance are more complex and likely involves several altered mechanisms, beyond just the deactivation of NHEJ. 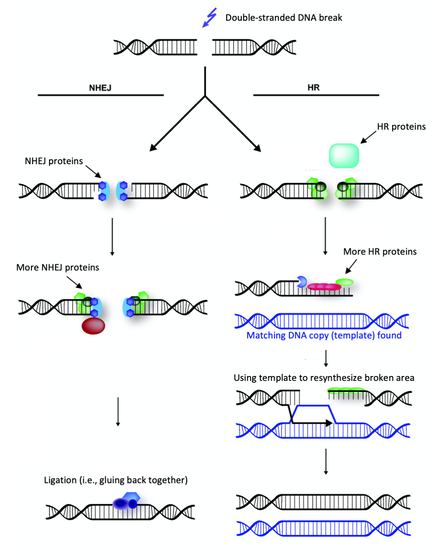 NHEJ vs HR. Source: modified from Wikimedia Commons. NHEJ vs HR. Source: modified from Wikimedia Commons. So, what is NHEJ and how could its deactivation help against ionizing radiation? NHEJ goes hand in hand with another DNA repair mechanism know as homologous recombination (HR). Both function in repairing double stranded DNA breaks—something that ionizing radiation is very good at inflicting. NHEJ fixes a break by directly ligating (i.e. gluing) the broken ends back together. HR repairs breaks by using a gene template (we have two copies of each gene—HR finds the copy and uses it as a template), which serves as a validating step to make sure the repair is accurate—something NHEJ doesn’t have. Organisms with smaller genomes (i.e., less DNA), such as fungi and bacteria, typically have a stronger predisposition for HR because they don’t possess large regions of genetic code repeats that could potentially result in inaccurate repair through HR. Romsdahl et al. propose two hypotheses to explain why selection for radiation resistance would favor deactivation of the NHEJ repair system. The first is that it reduces error-prone repair. Repair of DNA breaks by NHEJ is generally a precise and highly efficient process. However, radiation-induced DNA breaks usually produce chemically altered, or “dirty”, DNA ends, which cause imprecise repair by NHEJ. Further, during extreme radiation exposure, you’ll likely get a lot of DNA breaks, and as a result increase the probability of NHEJ gluing the wrong ends back together—this would be lethal. The second hypothesis is that deactivation of NHEJ reduces competition for repair by HR proteins. In yeast, DNA break repairs are first tackled by NHEJ. Breaks that are left unrepaired by NHEJ are then handed over to HR. Given that NHEJ is recruited to the break site before HR, NHEJ proteins bound to the break site can delay HR initiation. Increasing HR efficiency by eliminating competition from NHEJ proteins substantially improves survival following exposure to a high ionization radiation dose. In fact, HR function has been reported to increase threefold when it doesn’t have to compete with NHEJ [11] You might now be wondering if this discovery is something that could be directly applicable to us. Could we increase our resistance to ionizing radiation by deleting a few of our NHEJ genes? After all, rapidly advancing genome editing technologies are continuously pushing us towards the realm of human gene modifications. Unfortunately, like all things in biology, it’s not that simple. Depending on the organism and specific situation, shunting from NHEJ to HR for repair may not be feasible because there could be substantial survival consequences for disrupting NHEJ. Such a switch would be more attainable for species that predominantly rely on HR, like yeast, than for those that more heavily favor NHEJ, like us. And based on my own work in genetic engineering, a mutation that produces a beneficial effect in the gene of one animal, can have a completely different effect in the same gene of a different animal—adaptive outcomes are highly context dependent. However, each new discovery, like this one by Romsdahl et al., contributes to our growing understanding of the complex evolutionary mechanisms by which new adaptations are facilitated. One day we’ll get to the applicable part. By Shab Mohammadi
|
AuthorWant to learn about the latest discoveries in science? Here, I provide exciting monthly news updates on topics related to evolution. Archives
May 2021
Categories |
Proudly powered by Weebly
 RSS Feed
RSS Feed